-Search query
-Search result
Showing 1 - 50 of 1,585 items for (author: ise & t)

EMDB-43877: 
Human Amylin1 Receptor in Complex with Gs and human Calcitonin Gene-Related Peptide
Method: single particle / : Cao J, Belousoff MJ, Wootten DL, Sexton PM

PDB-9auc: 
Human Amylin1 Receptor in Complex with Gs and human Calcitonin Gene-Related Peptide
Method: single particle / : Cao J, Belousoff MJ, Wootten DL, Sexton PM

EMDB-36467: 
Cryo-EM structure of the head region of full-length ERGIC-53 with MCFD2 (form A)
Method: single particle / : Watanabe S, Inaba K

EMDB-36468: 
Cryo-EM structures of the head region of full-length ERGIC-53 with MCFD2 (form B)
Method: single particle / : Watanabe S, Inaba K

EMDB-36469: 
Cryo-EM structures of the head region of full-length ERGIC-53 with MCFD2 (Substate A)
Method: single particle / : Watanabe S, Inaba K

EMDB-36470: 
Cryo-EM structure of the head region of full-length ERGIC-53 with MCFD2 (Substate B)
Method: single particle / : Watanabe S, Inaba K

EMDB-36471: 
Cryo-EM structure of the head region of full-length ERGIC-53 with MCFD2 (Substate C)
Method: single particle / : Watanabe S, Inaba K

EMDB-36472: 
Cryo-EM structure of the head region of full-length ERGIC-53 with MCFD2 (Substate D)
Method: single particle / : Watanabe S, Inaba K

EMDB-36479: 
Cryo-EM structure of full-length ERGIC-53 with MCFD2
Method: single particle / : Watanabe S, Inaba K

EMDB-36482: 
cryoEM structure of ERGIC-53 deltaH34 mutant with MCFD2
Method: single particle / : Watanabe S, Inaba K

PDB-8jp4: 
Cryo-EM structure of the head region of full-length ERGIC-53 with MCFD2 (form A)
Method: single particle / : Watanabe S, Inaba K

PDB-8jp5: 
Cryo-EM structures of the head region of full-length ERGIC-53 with MCFD2 (form B)
Method: single particle / : Watanabe S, Inaba K

PDB-8jp6: 
Cryo-EM structures of the head region of full-length ERGIC-53 with MCFD2 (Substate A)
Method: single particle / : Watanabe S, Inaba K

PDB-8jp7: 
Cryo-EM structure of the head region of full-length ERGIC-53 with MCFD2 (Substate B)
Method: single particle / : Watanabe S, Inaba K

PDB-8jp8: 
Cryo-EM structure of the head region of full-length ERGIC-53 with MCFD2 (Substate C)
Method: single particle / : Watanabe S, Inaba K

PDB-8jp9: 
Cryo-EM structure of the head region of full-length ERGIC-53 with MCFD2 (Substate D)
Method: single particle / : Watanabe S, Inaba K

PDB-8jpg: 
Cryo-EM structure of full-length ERGIC-53 with MCFD2
Method: single particle / : Watanabe S, Inaba K
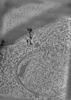
EMDB-38295: 
SPACEtomo example tomograms of Baker's Yeast
Method: electron tomography / : Eisenstein F, Danev R

EMDB-43736: 
Umb1 umbrella toxin particle
Method: single particle / : Park YJ, Zhao Q, Seattle Structural Genomics Center for Infectious Disease (SSGCID), DiMaio F, Mougous JD, Veesler D

EMDB-43737: 
Umb1 umbrella toxin particle (local refinement of UmbB1 bound ALF of UmbC1 and UmbA1)
Method: single particle / : Park YJ, Zhao Q, Seattle Structural Genomics Center for Infectious Disease (SSGCID), DiMaio F, Mougous JD, Veesler D

PDB-8w20: 
Umb1 umbrella toxin particle
Method: single particle / : Park YJ, Zhao Q, Seattle Structural Genomics Center for Infectious Disease (SSGCID), DiMaio F, Mougous JD, Veesler D

PDB-8w22: 
Umb1 umbrella toxin particle (local refinement of UmbB1 bound ALF of UmbC1 and UmbA1)
Method: single particle / : Park YJ, Zhao Q, Seattle Structural Genomics Center for Infectious Disease (SSGCID), DiMaio F, Mougous JD, Veesler D

EMDB-43889: 
Chlamydomonas reinhardtii mastigoneme (constituent map 1)
Method: single particle / : Dai J, Ma M, Zhang R, Brown A

EMDB-43890: 
Chlamydomonas reinhardtii mastigoneme (constituent map 2)
Method: single particle / : Dai J, Ma M, Zhang R, Brown A

EMDB-43891: 
Chlamydomonas reinhardtii mastigoneme (constituent map 3)
Method: single particle / : Dai J, Ma M, Zhang R, Brown A

EMDB-43892: 
Composite cryo-EM map of the Chlamydomonas reinhardtii mastigoneme
Method: single particle / : Dai J, Ma M, Zhang R, Brown A

PDB-9b4h: 
Chlamydomonas reinhardtii mastigoneme filament
Method: single particle / : Dai J, Ma M, Zhang R, Brown A

EMDB-19406: 
Structure of the human DDB1-DDA1-DCAF15 E3 ubiquitin ligase bound to compound furan 12
Method: single particle / : Shilliday F, Lucas SCC, Richter M, Michaelides IN, Fusani L

EMDB-19407: 
Structure of the human DDB1-DDA1-DCAF15 E3 ubiquitin ligase bound to compound furan 24
Method: single particle / : Shilliday F, Lucas SCC, Richter M, Michaelides IN, Fusani L

PDB-8rox: 
Structure of the human DDB1-DDA1-DCAF15 E3 ubiquitin ligase bound to compound furan 12
Method: single particle / : Shilliday F, Lucas SCC, Richter M, Michaelides IN, Fusani L

PDB-8roy: 
Structure of the human DDB1-DDA1-DCAF15 E3 ubiquitin ligase bound to compound furan 24
Method: single particle / : Shilliday F, Lucas SCC, Richter M, Michaelides IN, Fusani L

EMDB-41907: 
Computationally Designed, Expandable O4 Octahedral Handshake Nanocage
Method: single particle / : Weidle C, Borst A
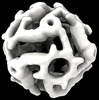
EMDB-42031: 
Computational Designed Nanocage O43_129_+8
Method: single particle / : Weidle C, Kibler RD

EMDB-43658: 
SARS-CoV-2 S (C.37 Lambda variant) plus S309, S2L20, and S2X303 Fabs
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-43659: 
SARS-CoV-2 S NTD (C.37 Lambda variant) plus S2L20 and S2X303 Fabs, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-43660: 
SARS-CoV-2 S RBD (C.37 Lambda variant) plus S309 Fab, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-8vye: 
SARS-CoV-2 S (C.37 Lambda variant) plus S309, S2L20, and S2X303 Fabs
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-8vyf: 
SARS-CoV-2 S NTD (C.37 Lambda variant) plus S2L20 and S2X303 Fabs, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-8vyg: 
SARS-CoV-2 S RBD (C.37 Lambda variant) plus S309 Fab, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-43318: 
Twistless helix 12 repeat ring design R12B
Method: single particle / : Calise SJ, Kollman JM

EMDB-42977: 
Cryo-EM structure of singly-bound SNF2h-nucleosome complex with SNF2h at inactive SHL2 (conformation 1)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-43000: 
Cryo-EM structure of SNF2h-nucleosome complex (consensus structure)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-43001: 
Cryo-EM structure of SNF2h-nucleosome complex (single-bound structure)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-43002: 
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-43003: 
Cryo-EM structure of singly-bound SNF2h-nucleosome complex with SNF2h at inactive SHL2 (conformation 2)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-43004: 
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex (conformation 1)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-43005: 
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex (conformation 2)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

PDB-8v4y: 
Cryo-EM structure of singly-bound SNF2h-nucleosome complex with SNF2h at inactive SHL2 (conformation 1)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

PDB-8v6v: 
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

PDB-8v7l: 
Cryo-EM structure of singly-bound SNF2h-nucleosome complex with SNF2h at inactive SHL2 (conformation 2)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
